This section provides information about all functions and variables related to the calculation of the partition function and base pair probabilities. More...
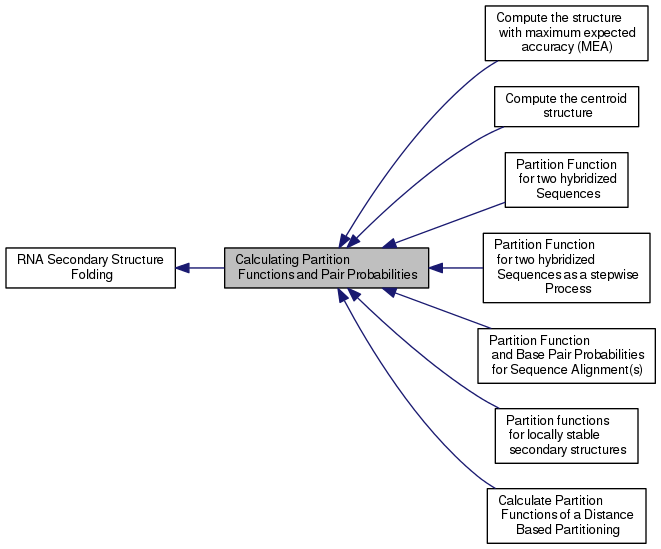
 Collaboration diagram for Calculating Partition Functions and Pair Probabilities:
Collaboration diagram for Calculating Partition Functions and Pair Probabilities:Modules | |
| Compute the structure with maximum expected accuracy (MEA) | |
| Compute the centroid structure | |
| Partition Function for two hybridized Sequences | |
| Partition Function Cofolding. | |
| Partition Function for two hybridized Sequences as a stepwise Process | |
| Partition Function Cofolding as a stepwise process. | |
| Partition Function and Base Pair Probabilities for Sequence Alignment(s) | |
| Partition functions for locally stable secondary structures | |
| Calculate Partition Functions of a Distance Based Partitioning | |
| Compute the partition function and stochastically sample secondary structures for a partitioning of the secondary structure space according to the base pair distance to two fixed reference structures. | |
Files | |
| file | part_func.h |
| Partition function of single RNA sequences. | |
Functions | |
| float | pf_fold_par (const char *sequence, char *structure, pf_paramT *parameters, int calculate_bppm, int is_constrained, int is_circular) |
Compute the partition function 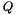 for a given RNA sequence. More... for a given RNA sequence. More... | |
| float | pf_fold (const char *sequence, char *structure) |
Compute the partition function 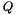 of an RNA sequence. More... of an RNA sequence. More... | |
| float | pf_circ_fold (const char *sequence, char *structure) |
| Compute the partition function of a circular RNA sequence. More... | |
| void | free_pf_arrays (void) |
| Free arrays for the partition function recursions. More... | |
| void | update_pf_params (int length) |
| Recalculate energy parameters. More... | |
| void | update_pf_params_par (int length, pf_paramT *parameters) |
| Recalculate energy parameters. | |
| double * | export_bppm (void) |
| Get a pointer to the base pair probability arrayAccessing the base pair probabilities for a pair (i,j) is achieved by. More... | |
| void | assign_plist_from_pr (plist **pl, double *probs, int length, double cutoff) |
| Create a plist from a probability matrix. More... | |
| int | get_pf_arrays (short **S_p, short **S1_p, char **ptype_p, double **qb_p, double **qm_p, double **q1k_p, double **qln_p) |
| Get the pointers to (almost) all relavant computation arrays used in partition function computation. More... | |
| double | mean_bp_distance (int length) |
| Get the mean base pair distance of the last partition function computation. More... | |
| double | mean_bp_distance_pr (int length, double *pr) |
| Get the mean base pair distance in the thermodynamic ensemble. More... | |
Detailed Description
This section provides information about all functions and variables related to the calculation of the partition function and base pair probabilities.
Instead of the minimum free energy structure the partition function of all possible structures and from that the pairing probability for every possible pair can be calculated, using a dynamic programming algorithm as described in[10].
Function Documentation
| float pf_fold_par | ( | const char * | sequence, |
| char * | structure, | ||
| pf_paramT * | parameters, | ||
| int | calculate_bppm, | ||
| int | is_constrained, | ||
| int | is_circular | ||
| ) |
Compute the partition function 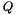 for a given RNA sequence.
for a given RNA sequence.
If structure is not a NULL pointer on input, it contains on return a string consisting of the letters " . , | { } ( ) " denoting bases that are essentially unpaired, weakly paired, strongly paired without preference, weakly upstream (downstream) paired, or strongly up- (down-)stream paired bases, respectively. If fold_constrained is not 0, the structure string is interpreted on input as a list of constraints for the folding. The character "x" marks bases that must be unpaired, matching brackets " ( ) " denote base pairs, all other characters are ignored. Any pairs conflicting with the constraint will be forbidden. This is usually sufficient to ensure the constraints are honored. If the parameter calculate_bppm is set to 0 base pairing probabilities will not be computed (saving CPU time), otherwise after calculations took place pr will contain the probability that bases i and j pair.
- Note
- The global array pr is deprecated and the user who wants the calculated base pair probabilities for further computations is advised to use the function export_bppm()
- Postcondition
- After successful run the hidden folding matrices are filled with the appropriate Boltzmann factors. Depending on whether the global variable do_backtrack was set the base pair probabilities are already computed and may be accessed for further usage via the export_bppm() function. A call of free_pf_arrays() will free all memory allocated by this function. Successive calls will first free previously allocated memory before starting the computation.
- See Also
- pf_fold(), pf_circ_fold(), bppm_to_structure(), export_bppm(), get_boltzmann_factors(), free_pf_arrays()
- Parameters
-
[in] sequence The RNA sequence input [in,out] structure A pointer to a char array where a base pair probability information can be stored in a pseudo-dot-bracket notation (may be NULL, too) [in] parameters Data structure containing the precalculated Boltzmann factors [in] calculate_bppm Switch to Base pair probability calculations on/off (0==off) [in] is_constrained Switch to indicate that a structure contraint is passed via the structure argument (0==off) [in] is_circular Switch to (de-)activate postprocessing steps in case RNA sequence is circular (0==off)
- Returns
- The Gibbs free energy of the ensemble (
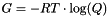 ) in kcal/mol
) in kcal/mol
| float pf_fold | ( | const char * | sequence, |
| char * | structure | ||
| ) |
Compute the partition function 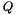 of an RNA sequence.
of an RNA sequence.
If structure is not a NULL pointer on input, it contains on return a string consisting of the letters " . , | { } ( ) " denoting bases that are essentially unpaired, weakly paired, strongly paired without preference, weakly upstream (downstream) paired, or strongly up- (down-)stream paired bases, respectively. If fold_constrained is not 0, the structure string is interpreted on input as a list of constraints for the folding. The character "x" marks bases that must be unpaired, matching brackets " ( ) " denote base pairs, all other characters are ignored. Any pairs conflicting with the constraint will be forbidden. This is usually sufficient to ensure the constraints are honored. If do_backtrack has been set to 0 base pairing probabilities will not be computed (saving CPU time), otherwise pr will contain the probability that bases i and j pair.
- Note
- The global array pr is deprecated and the user who wants the calculated base pair probabilities for further computations is advised to use the function export_bppm().
- OpenMP: This function is not entirely threadsafe. While the recursions are working on their own copies of data the model details for the recursions are determined from the global settings just before entering the recursions. Consider using pf_fold_par() for a really threadsafe implementation.
- Precondition
- This function takes its model details from the global variables provided in RNAlib
- Postcondition
- After successful run the hidden folding matrices are filled with the appropriate Boltzmann factors. Depending on whether the global variable do_backtrack was set the base pair probabilities are already computed and may be accessed for further usage via the export_bppm() function. A call of free_pf_arrays() will free all memory allocated by this function. Successive calls will first free previously allocated memory before starting the computation.
- Parameters
-
sequence The RNA sequence input structure A pointer to a char array where a base pair probability information can be stored in a pseudo-dot-bracket notation (may be NULL, too)
- Returns
- The Gibbs free energy of the ensemble (
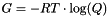 ) in kcal/mol
) in kcal/mol
| float pf_circ_fold | ( | const char * | sequence, |
| char * | structure | ||
| ) |
Compute the partition function of a circular RNA sequence.
- Note
- The global array pr is deprecated and the user who wants the calculated base pair probabilities for further computations is advised to use the function export_bppm().
- OpenMP: This function is not entirely threadsafe. While the recursions are working on their own copies of data the model details for the recursions are determined from the global settings just before entering the recursions. Consider using pf_fold_par() for a really threadsafe implementation.
- Precondition
- This function takes its model details from the global variables provided in RNAlib
- Postcondition
- After successful run the hidden folding matrices are filled with the appropriate Boltzmann factors. Depending on whether the global variable do_backtrack was set the base pair probabilities are already computed and may be accessed for further usage via the export_bppm() function. A call of free_pf_arrays() will free all memory allocated by this function. Successive calls will first free previously allocated memory before starting the computation.
- See Also
- pf_fold_par(), pf_fold()
- Parameters
-
[in] sequence The RNA sequence input [in,out] structure A pointer to a char array where a base pair probability information can be stored in a pseudo-dot-bracket notation (may be NULL, too)
- Returns
- The Gibbs free energy of the ensemble (
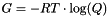 ) in kcal/mol
) in kcal/mol
| void free_pf_arrays | ( | void | ) |
Free arrays for the partition function recursions.
Call this function if you want to free all allocated memory associated with the partition function forward recursion.
- Note
- Successive calls of pf_fold(), pf_circ_fold() already check if they should free any memory from a previous run.
-
OpenMP notice:
This function should be called before leaving a thread in order to avoid leaking memory
- Postcondition
- All memory allocated by pf_fold_par(), pf_fold() or pf_circ_fold() will be free'd
- See Also
- pf_fold_par(), pf_fold(), pf_circ_fold()
| void update_pf_params | ( | int | length | ) |
Recalculate energy parameters.
Call this function to recalculate the pair matrix and energy parameters after a change in folding parameters like temperature
| double* export_bppm | ( | void | ) |
Get a pointer to the base pair probability arrayAccessing the base pair probabilities for a pair (i,j) is achieved by.
- Precondition
- Call pf_fold_par(), pf_fold() or pf_circ_fold() first to fill the base pair probability array
- See Also
- pf_fold(), pf_circ_fold(), get_iindx()
- Returns
- A pointer to the base pair probability array
| void assign_plist_from_pr | ( | plist ** | pl, |
| double * | probs, | ||
| int | length, | ||
| double | cutoff | ||
| ) |
Create a plist from a probability matrix.
The probability matrix given is parsed and all pair probabilities above the given threshold are used to create an entry in the plist
The end of the plist is marked by sequence positions i as well as j equal to 0. This condition should be used to stop looping over its entries
- Note
- This function is threadsafe
- Parameters
-
[out] pl A pointer to the plist that is to be created [in] probs The probability matrix used for creting the plist [in] length The length of the RNA sequence [in] cutoff The cutoff value
| int get_pf_arrays | ( | short ** | S_p, |
| short ** | S1_p, | ||
| char ** | ptype_p, | ||
| double ** | qb_p, | ||
| double ** | qm_p, | ||
| double ** | q1k_p, | ||
| double ** | qln_p | ||
| ) |
Get the pointers to (almost) all relavant computation arrays used in partition function computation.
- Precondition
- In order to assign meaningful pointers, you have to call pf_fold_par() or pf_fold() first!
- See Also
- pf_fold_par(), pf_fold(), pf_circ_fold()
- Parameters
-
[out] S_p A pointer to the 'S' array (integer representation of nucleotides) [out] S1_p A pointer to the 'S1' array (2nd integer representation of nucleotides) [out] ptype_p A pointer to the pair type matrix [out] qb_p A pointer to the QB matrix [out] qm_p A pointer to the QM matrix [out] q1k_p A pointer to the 5' slice of the Q matrix ( 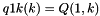 )
) [out] qln_p A pointer to the 3' slice of the Q matrix ( 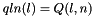 )
)
- Returns
- Non Zero if everything went fine, 0 otherwise
| double mean_bp_distance | ( | int | length | ) |
Get the mean base pair distance of the last partition function computation.
- Note
- To ensure thread-safety, use the function mean_bp_distance_pr() instead!
- See Also
- mean_bp_distance_pr()
- Parameters
-
length
- Returns
- mean base pair distance in thermodynamic ensemble
| double mean_bp_distance_pr | ( | int | length, |
| double * | pr | ||
| ) |
Get the mean base pair distance in the thermodynamic ensemble.
This is a threadsafe implementation of mean_bp_dist() !
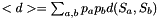
this can be computed from the pair probs 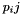 as
as
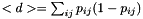
- Note
- This function is threadsafe
- Parameters
-
length The length of the sequence pr The matrix containing the base pair probabilities
- Returns
- The mean pair distance of the structure ensemble